 Loading
......
Loading
......
PseudoUridine and Its Derivatives
PseudoUridine (abbreviated by Ψ or Q) is an isomer of the nucleoside uridine (abbreviated by U) in which the uracil is bound with ribose through a carbon-carbon rather than a nitrogen-carbon glycosidic bond. PseudoUridine has been called the “fifth nucleotide” of RNA, due to its prevalence in rRNAs, tRNAs, and snRNAs and is the most abundant modification in total cellular RNA. PseudoUridine is formed via isomerization of uridine such that the ribose C1 binds to uracil C5 instead of N1, and the more accessible N1 can form additional hydrogen bonds, which leads to a more stable RNA secondary structure and increases the rigidity of RNA. Derivatives of pseudoUridine can be obtained by N1 and/or N3 functionalization on uracil, or by deoxidation of ribose, or protection of hydroxyl groups on ribose.
PseudoUridine Synthases
PseudoUridine synthases catalyze the isomerization of U to Ψ in RNA. These enzymes use both sequence and structural information from their RNA substrates to achieve this transformation. The Ψ synthases are divided into six different families that do not share significant global sequence similarity, and they are thought to be a result of divergent evolution. And structural studies show that all Ψ synthases share a core β-fold and a conserved catalytic domain. An active site aspartic acid (Asp) is conserved among all Ψ synthases, and it is absolutely essential for the catalytic activity of Ψ enzymes.
The pseudoUridine synthases isomerize U in RNA to pseudoUridine Ψ, and the mechanism that they follow has long been a question of interest. The possible mechanism, “glycal mechanism”, has been reported that based on the observed epimerization at C2′ of the ribose ring of 5-fluorouridine in RNA, which might be achieved through deprotonation of C2′ and then reprotonation of the resulting glycal intermediate from the opposite face.
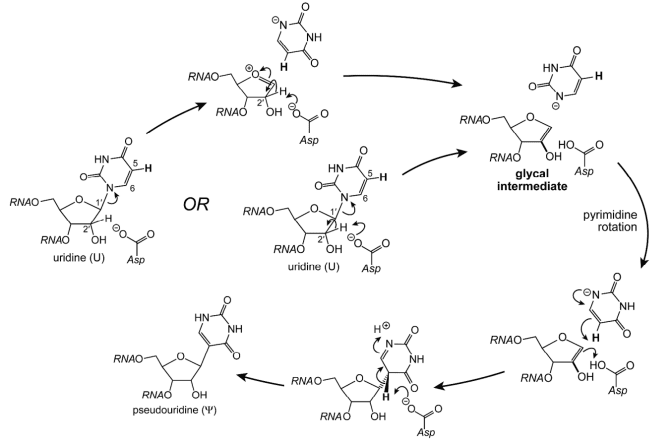
Applications
The nucleoside modification is an effective approach to enhance stability and translational capacity of mRNA while diminishing its immunogenicity in vivo. PseudoUridine can be used to improve properties of mRNA and to make such mRNA a promising tool for both gene replacement and vaccination. Incorporating pseudoUridine into mRNA not only suppresses RNA-mediated immune activation in vitro and in vivo, but also enhances the translational capacity of the RNA. These characteristics and the ease of generating such an RNA by in vitro transcription make ψ-containing mRNA a unique tool for expression of any protein in vitro and in vivo.
Pseudouridylation is thought to promote stability and conformation of mitochondrial and nuclear tRNAs. Mutations in pseudoUridine synthase 1 (PUS1) differentially impair nuclear and mitochondrial tRNA maturation causing mitochondrial myopathy, lactic acidosis, and sideroblastic anemia (MLASA). And pseudoUridines have been recognized as regulators of viral latency processes in human immunodeficiency virus (HIV) infections.
Derivatives
Post-transcriptional modifications are an integral part of naturally occurring functional RNAs. One of the most commonly occurring modifications in RNA is pseudoUridine Ψ. Apart from occurring in tRNAs, Ψ is commonly observed in ribosomal RNAs (rRNAs). RNAs also contain modifications, which are derivatives of Ψ. These include 1-methylpseudoUridine (m1 Ψ), 3-methylpseudoUridine (m3Ψ), 2′-O-methylpseudoUridine (Ψm), and a hypermodified residue 1-methyl-3- (3-amino-3-carboxypropyl) pseudoUridine (m1acp3Ψ).
1-methylpseudoUridine (m1 Ψ)
1-MethylpseudoUridine was first observed in the tRNAs of archaebacteria in which they were found to occur in the TΨC loop in place of ribothymidine. The base pairing properties and structure of this residue is very similar to uridine or ribothymidine. It may also confer an advantage in the thermal stability of the elbow region in tRNAs due to its enhanced stacking properties. Interestingly, therapeutic mRNAs with 1-methylpseudoUridine modification show markedly increased expressivity upon transfection into cell lines and reduced innate immunogenicity compared to pseudoUridine modified mRNAs.
3-methylpseudoUridine (m3Ψ)
3-MethylpseudoUridine was first identified in a ribosomal RNA in 1996 although its synthesis has been previously reported in 1982. Although the N3 position of this nucleoside is incapable of forming hydrogen bonds, it can still interact with adenine through the N1 position if it adopts the syn conformation. There is little idea why 3- methylpseudoUridine is so strongly conserved at position 1915 in helix 69 in prokaryotic ribosomal RNA. Mutation of m3 Ψ at the 1915 position has been found to impair translation fidelity and subunit association.
2′-O-methylpseudoUridine (Ψm)
The 2′-O-methylated derivative of pseudoUridine has been observed in tRNAs of archaea and eukaryotes, in the 18+26S rRNA of eukaryotes, and in snRNAs in eukaryotes. In general, the 2′-O-methylation of the ribose stabilizes the C3′- endo conformation and makes the base pairing more thermodynamically stable in RNA. It can also disrupt the participation of the 2′-OH group in tertiary interactions. The steric effect of the methyl group can also influence RNA−protein interactions. CD and NMR studies suggested that although Ψm preferred the syn conformation predominantly, the preference was less than that of Ψ. In contrast to most other 2′-O-methylated nucleosides, Ψm was observed to slightly prefer the C2′-endo conformation over the C3′-endo conformation.
1-methyl-3- (3-amino-3-carboxypropyl) pseudoUridine (m1acp3Ψ).
The hypermodified residue 1-methyl-3-(3-amino-3-carboxypropyl) pseudoUridine was first isolated from the RNA of cells of an established line of cells of Chinese hamster in 1974 and was localized in 18S rRNAs. It was shown that both the methyl and the α-aminobutyrate moieties are derived from methionine. Later, its presence was confirmed in HeLa 18S and yeast 17S rRNA. Very recently, this modification has been precisely located at position 1248 of the 18S rRNA in the human 80S ribosome. It has been postulated that in correspondence with the well-known tendency of amino acid side chains to chelate divalent cations and form hydrogen bonds, the amino acid side-chain modifications of nucleosides such as m1 acp3 Ψ can carry out these functions.




